Kaggle Competition: BNP Paribas Cardif Claims Management
Contributed by Ablat, Bin Lin, Claudia Huang and Sung Pil Moon. They took NYC Data Science Academy 12 week full time Data Science Bootcamp program between Jan. 11th to April 1st, 2016.This post is based on their fourth class project - Machine Learning (due on the 8th week of the program).
1. Overview
As part of a Kaggle competition, we are challenged to help BNP Paribas cardif to accelerate their claims management process in order to provide a better service to its customers.
In today's world, everything is going faster and faster. When facing unexpected events, customers expect their insurer to support them as soon as possible when facing unexpected events. However, claims management may require different levels of check before a claim can be approved and a payment can be made. In this challenge, we are going to classify the claims into the following two categories:
- claims for which approval could be accelerated leading to faster payments
- claims for which additional information is required before approval
In this project, we have done:
- Exploratory data analysis
- Feature engineering
- Missing data imputation
- Building prediction models with various machine learning algorithms, including Logistic Regression, Decision Trees, Random Forest, Xgboost, Neural Network.
2. The Data set
2.1. Data Summary
The data provided by BNP Paribas cardif are:
- TRAIN.CSV: traing dataset
- TEST.CSV: test dataset
- SAMPLE_SUBMISSION.CSV: example of submission format
These are the basic description of the training dataset:
- Total: 111,432 observations (rows) x 133 features (columns)
- Data Types: float, integer, string
- Column "ID": the ID of each row, not being used as predictor
- Column "target": the target of each row, not being used as predictor
- Numbers of Text-based Predictors: 112
- Numbers of Numeric Predictors: 19
- Numbers of Columns with missing value: 119
- Numbers of columns highly correlated: There are 123 pairs with absolute correlation > 0.8; there are 63 pairs with absolute correlation > 0.9.
2.2. Exploratory Data Analysis
As the first step, we performed exploratory data analysis to better understand the data. These are the initial findings throughout the EDA process.
- Anonymized Data: All data (both categorical and continuous) is anonymized without any description
| ID | target | v1 | v2 | v3 | ... | v127 | v128 | v129 | v130 | v131 |
|---|---|---|---|---|---|---|---|---|---|---|
| 3 | 1 | 1.335739 | 8.727474 | C | ... | 3.113719 | 2.024285 | 0 | 0.636365 | 2.857144 |
| 4 | 1 | NaN | NaN | C | ... | NaN | 1.957825 | 0 | NaN | NaN |
| 5 | 1 | 0.943877 | 5.310079 | C | ... | 3.922193 | 1.120468 | 2 | 0.883118 | 1.176472 |
| 6 | 1 | 0.797415 | 8.304757 | C | ... | 2.954381 | 1.990847 | 1 | 1.677108 | 1.034483 |
| 8 | 1 | NaN | NaN | C | ... | NaN | NaN | 0 | NaN | NaN |
Table 1: Data First Peek
- Too Many Missing Value: Approximately 40% of data is missing as you can see in the Figure 1.
- High correlations: As shown in the correlation matrix plot (Figure 2), some variables are highly correlated (not just 1-to-many but also many-to-many). Correlation values are between -1 and 1. Red color means positive correlation while blue color means negative correlation.
3. Data Pre-Processing
Data pre-processing is an important step in our before we can even start building our models. Our data pre-processing includes, data cleaning, transformation, feature selection.
3.1. Data Cleaning
3.1.1. Treatment of Integer Variables with Low Numbers of Unique Values
There are four of the integer variables with low numbers of unique values: v38, v62, v72, v129. These will be treated as categorical variables. These variables would be factorized using binary dummies.
| index | _a_variable | _b_data_type | _c_cardinality | _d_missings | _e_sample_values |
|---|---|---|---|---|---|
| 39 | v38 | int64 | 12 | 0 | [0, 4] |
| 63 | v62 | int64 | 8 | 0 | [1, 2] |
| 73 | v72 | int64 | 13 | 0 | [1, 2] |
| 130 | v129 | int64 | 10 | 0 | [0, 2] |
Table 2: Integer Variables with Low Cardinality
3.1.2. Imputation
Due to the high missingess in the dataset, we have sent a lot of time on data exploratory and figuring out the best imputation methods. The following are the methods we have tried.
- Numeric Imputation with Mean: We chose to imputed the numeric variables with mean to start with our data training.
- Numeric Imputation with Interpolate Linear
- Numeric Imputation with -999: impute with extreme numbers
- Categorical Imputation with "NA": Dropping all missing value in the categorical values does not help increasing the higher predictability. Some missingness itself can provide information, such as for the question like "How long have been in a marriage?", NA means 0 years of marriage.
- Prediction Model: which uses supervised algorithms (Linear regression, KNN, Tree-based, etc) to predict the variables with missingness based on other variables. When we tried this method with KNN and Decision Tree, it took forever to run and never been able to finish. Due to tight timeline, we ended up giving up on this method.
As some experiment, we go with imputing numeric variables with -999, and categorical variables with text "NA" as these two options give us a better results.
3.2. Data Transformation
Many machine learning tool, including Python, will only accept numbers as input. This was a problem as Python was being used in our project. Fortunately, the Pandas package has get_dummies() function, which converts categorical variable into dummy/indicator variables.
Sample code: convert categorical data into binary dummy variables
# convert text-based columns to dummies for var_name in cate_variables: dummies = pd.get_dummies(df[var_name], prefix=var_name) # Drop the current variable, concat/append the dummy dataframe to the dataframe. df_new = pd.concat([df_new.drop(var_name, 1), dummies.iloc[:,1:]], axis = 1)
3.3. Feature Selection
3.3.1. Remove Highly Correlated Variables
As mentioned earlier, there are 63 variable pairs with absolute correlation > 0.9. Between the two variables in a pair, the one with higher missingness would be removed from the training model. As it may not needed by tree-based algorithms, it helps improve the performance by having fewer features.
| index | _var1 | _var2 | _var_corr | var1_na | var2_na |
|---|---|---|---|---|---|
| 0 | v12 | v10 | 0.912 | 49851 | 84 |
| 1 | v25 | v8 | 0.943 | 49840 | 48619 |
| 2 | v32 | v15 | 0.908 | 48619 | 49836 |
| 3 | v40 | v34 | -0.903 | 49832 | 111 |
| 4 | v41 | v29 | 0.904 | 49832 | 49832 |
| 5 | v43 | v26 | 0.903 | 49832 | 49832 |
| ... | ... | ... | ... | ... | ... |
| 56 | v115 | v69 | -0.994 | 49843 | 49895 |
| 57 | v116 | v43 | 0.978 | 49832 | 49836 |
| 58 | v118 | v97 | 0.962 | 49843 | 49843 |
| 59 | v121 | v33 | 0.949 | 48654 | 49832 |
| 60 | v121 | v83 | 0.966 | 48654 | 49832 |
| 61 | v128 | v108 | 0.957 | 49832 | 48624 |
| 62 | v128 | v109 | 0.903 | 49832 | 48624 |
Table 3: Highly Correlated Variables
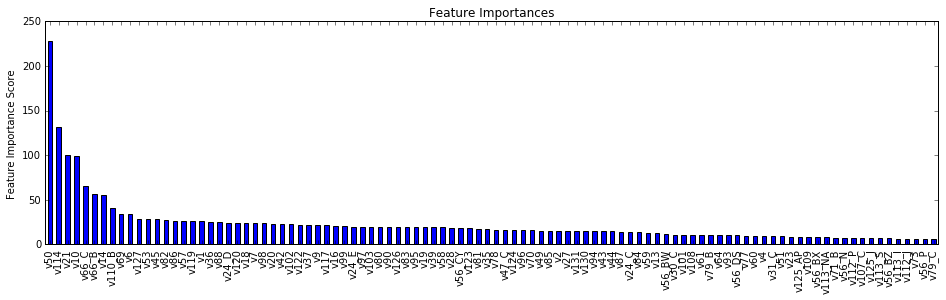
3.3.2. Tree-based Feature Selection
Many tree-based methods provides feature importance, which can be used to discard the irrelevant features. We used the ExtraTrees classifier and coupled it with the "SelectFromModel" in the scikit-learn package "feature_selection". SelectFromModel can be used along with any model that has a coef_ or feature_importances_ attribute after fitting. It also allows us to set the threshold for feature selection. Features whose importance is greater or equal are kept while the others are discarded. We picked 0.0003 as the threshold and got the 53 features returned:
['v1','v10','v101','v102','v105','v107','v108','v109','v110','v112', 'v113','v117','v119','v123','v124','v125','v129','v131','v14','v16', 'v2','v21','v22','v23','v24','v3','v30','v31','v34','v36','v38','v45', 'v47','v5','v50','v51','v52','v56','v58','v62','v64','v66','v69','v70', 'v71','v72','v74','v75','v78','v79','v82','v85','v87','v9','v91','v98']
def select_features_from_model(X_train, target_train, threshold): # Create a ExtraTreesClassifier with initial parameters model = ExtraTreesClassifier( n_estimators = 200, # Number of trees max_features = 0.8, # Number of features for each tree max_depth = 10, # Depth of the tree min_samples_split = 4, # Minimum number of samples required to split min_samples_leaf = 2, # Minimum number of samples in a leaf min_weight_fraction_leaf = 0, # Minimum weighted fraction of the input samples required to be at a leaf node. criterion = 'gini', # Use gini, not going to tune it random_state = 27, n_jobs = -1) model_fit = model.fit(X_train, target_train) model_select = SelectFromModel(model, threshold, prefit=True) model_select.transform(X_train) features_selected = X_train.columns[model_select.get_support()] features_dropped = X_train.columns[~model_select.get_support()] return (features_selected, features_dropped)
3.3.3. Univariate Feature Selection
We also tried univariate feature selection method which selects the best features based on univariate statistical tests. Again from the same package, we used the "SelectPercentile" method to perform a ANOVA F-test for classification tasks to retrieve the best features. Know the important features from the ExtraTree classifier were around 50, which was approximately 50% of the features after removing some highly correlated variables. we were interested in the top 50% of the features. We retrieved 47 features this time.
['v103','v105','v108','v109','v110','v113','v117','v119','v122', 'v123','v124','v129','v13','v130','v14','v16','v21','v23','v24', 'v28','v31','v36','v38','v45','v47','v5','v51','v58','v62','v66', 'v68','v69','v70','v72','v73','v74','v78','v79','v80','v82','v83', 'v84','v85','v86','v87','v9','v98']
def select_features_univariate(X_train, target_train, percent): model_select = SelectPercentile(f_classif, percent) model_select.fit_transform(X_train, target_train) features_selected = X_train.columns[model_select.get_support()] features_dropped = X_train.columns[~model_select.get_support()] return (features_selected, features_dropped)
4. Model Building
4.1. Machine Learning Methods
In this competition, we have tried a number of different machine learning methods. Below is a quick review of the methods with a general description, their advantages and disadvantages.
| Classifier | Description | Advantages | Disadvantages |
|---|---|---|---|
| Logistic Regression | Use regression to solve binary classification problems by fitting a sigmoid funciton to calculate the probabilities |
|
|
| Random Forest | Uses the random subsets of observations and features from the training data to create a number of decision trees; use averaging or voting to improve the predictive accuracy and control over-fitting. |
|
|
| Extra Trees Classifier | Is similar to Random Forest, fits a number of randomized decision trees on random subsets of observations and features; when choosing variables at a split, samples are drawn from the entire training set; splits are chosen completely at random from the range of values in the sample at each split. |
|
|
| GBM (Gradient Boosting Model) | Trains trees sequentially, uses the information from the previous trees, then combined the whole set and averaging to provide a higher prediction; it uses the entire features, and does not ignore 'weak' learners. |
|
|
| XGBoost (Extreme Gradient Boost) | Is an advanced implementation of gradient boosting algorithm; it has definitely boosting capabilities and additional components overcoming the weakness of GBM. |
|
|
Table 4: Summary of Machine Learning Methods
4.2. Procedure
After data cleaning, data imputation, and features selections, we started building our models. There were a few common steps during building our classification models:
- Training data sampling: We first trained our model with the full training set. However for some algorithms, it took very long time to tune the parameters and fit the model. Therefore about 30%-40% of the training data were used.
- Model fitting for the first try: This gave us a first taste of how the models performed at the first try with some initial parameters.
- Parameter tuning: For parameter tuning, we did grid-search using the GridSearchCV package from skitlearn. Since doing a complete grid-search is time consuming on a big dataset on a local computer, we used coarse grid search first. After identifying a "better" region on the gird, a finer gird search on that region can be conducted. Also for some of the algorithms (e.g. ExtraTreeClassifier and Xgboost) we broke out the parameters into groups and tuned them group by group.
- Fit the best estimator with the full training set: After we found the best estimator, we refitted the model with full training data to get a better model. Usually more data is better.
- Prediction with test data: as the final step, the test data is passed to the model to get the prediction probability.
4.3. Parameters Tuning
Due to the time constraint, we didn't get a chance to finish building models for all the methods. The three models we were able to finish are: Logistic Regression, ExtraTreeClassifier, XGBoost. Below are details of our parameters tuning details for these models.
4.3.1. ExtraTreesClassifier
We first built a ExtraTreesClassifier model and initialized the parameters with some reasonable values. We fit the model with our training samples. It returned score (using Log Loss scoring metric) : 0.4432684.
############ Create a ExtraTreesClassifier with initial parameters ######## model = ExtraTreesClassifier( n_estimators = 100, # Number of trees max_features = 0.8, # Number of features for each tree max_depth = 10, # Depth of the tree min_samples_split = 2, # Minimum number of samples required to split min_samples_leaf = 1, # Minimum number of samples in a leaf min_weight_fraction_leaf = 0, # Minimum weighted fraction of the input samples required to be at a leaf node. criterion = 'gini', # Use gini, not going to tune it random_state = 27, n_jobs = -1)
Next, we were going to use GridSearchCV to tune the parameters. For ExtraTreesClassifier, we were tuning the following five parameters. Note that we didn't tune n_estimators (number of trees) yet as larger number of trees means slower process. We left it to the end to find a reasonable value n_estimators.
Tuning parameters:
- n_estimators (not tuned at the beginning)
- max_features
- max_depth
- min_samples_split
- min_samples_leaf
We started coarse parameter search based on the initial model. We assigned three values for each parameter (except n_estimators), which gave us 81 combinations.
####### Coarse Tune Parameters ####### para_grid = [{ 'max_features': [0.6, 0.75, 0.9], # Number of features for each tree 'max_depth': [5, 15, 25], # Depth of the tree 'min_samples_split': [5, 10, 50], # Minimum number of samples required to split an internal node 'min_samples_leaf': [5, 10, 50] # Minimum number of samples in a leaf }] start = datetime.datetime.now() para_search = GridSearchCV(model, para_grid, scoring = 'log_loss', cv = 5, n_jobs = 4).fit(X_train, target_train) end = datetime.datetime.now() print "model training time: {}".format(end - start)
The result of coarse grid search gave the best combination: {'max_features': 0.9, 'min_samples_split': 5, 'max_depth': 25, 'min_samples_leaf': 50}, and the best Score 0.4740551.
After we found the best combination from coarse search, we performed a finer search in a narrower search range around the values we found previously.
####### Now Fine Tune Parameters ####### para_grid = [{ 'max_features': [0.85, 0.9, 0.95],# Number of features for each tree 'max_depth': [20, 25, 30], # Depth of the tree 'min_samples_split': [3, 5, 7], # Minimum number of samples required to split 'min_samples_leaf': [45, 50, 55] # Minimum number of samples in a leaf }]
The grid search gave the best combination: {'max_features': 0.85, 'min_samples_split': 3, 'max_depth': 25, 'min_samples_leaf': 45}, the score was improved to: 0.4736519. We can see that max_depth remains 25 from this round. We used 25 as the final value for max_depth (we could have continued to tune around 25 if we had more time). However the other three parameters are all changed and thus need further search.
We repeated the same steps to fine tunes the rest of the parameters. In the last round, we searched n_estimators in the range of [100, 300, 500, 700]. The grid search showed that 700 gave better score. Given that 700 was already a big number of trees, we stopped here without going further testing a bigger number which would make it a very long time to train our data. The table below shows the ranges we used to perform the gird search for different values as well as result for each round.
| n_estimators | max_features | max_depth | min_samples_split | min_samples_leaf | best combination | score |
|---|---|---|---|---|---|---|
| 100 | [0.6, 0.75, 0.9] | [5, 15, 25] | [5, 10, 50] | [5, 10, 50] |
|
0.4740551 |
| 100 | [0.85, 0.9, 0.95] | [20, 25, 30] | [3, 5, 7] | [45, 50, 55] |
|
0.4736519 |
| 100 | [0.83, 0.85, 0.87] | [25] | [2, 3, 4] | [42, 45, 48] |
|
0.4735396 |
| 100 | [0.86, 0.87, 0.88] | [25] | [2] | [45] |
|
0.4735396 |
| [100, 300, 500, 700] | [0.87] | [25] | [2] | [45] |
|
0.4733141 |
Table 5: ExtraTrees Parameters Search
We then fitted the model with the full training data and then got a much better training score 0.468914.
4.3.2. XGBoost
During building XGBoost model, we also built a base model and with some reasonable values for the parameters. We fit the model with our training samples. It returned score (using Log Loss scoring metric) : 0.434604.
######### Set initial values for the model ###### xgb_model = XGBClassifier( learning_rate =0.1, n_estimators=1000, max_depth=5, min_child_weight=1, gamma=0, subsample=0.8, colsample_bytree=0.8, objective= 'binary:logistic', nthread=8, scale_pos_weight=1, seed=27)
XGBClassifier(base_score=0.5, colsample_bylevel=1, colsample_bytree=0.8,
gamma=0, learning_rate=0.1, max_delta_step=0, max_depth=5,
min_child_weight=1, missing=None, n_estimators=97, nthread=8,
objective='binary:logistic', reg_alpha=0, reg_lambda=1,
scale_pos_weight=1, seed=27, silent=True, subsample=0.8)
For XGB, we were tuning the following seven parameters.
- learning_rate: learning rate (not going to be tune at the beginning)
- max_depth: depth of the trees
- min_child_weight: minimum number of observations in a node
- gamma: minimum loss reduction to make a split
- subsample: number of selected rows for training sample
- colsample_bytree: number of features for training sample
- reg_alpha: L1 regularization term on weight (analogous to Lasso regression)
One really useful benefit of XGB is the built-in tree-pruning. It would stop generating more trees at certain point when more trees do not reduce the errors. Therefore we can estimate n_estimators from the initial fit without performing a search for it. Also we used a fixed learning rate of 0.1 to allow faster performance as lower learning rate causes longer time to fit the model. We left it to the end to tune it.
Since the number of combination is big with seven parameters, we took the step by step approach.
We tuned the max_depth and min_child_weight parameters first as they will have the highest impact on model outcome. We started with wider ranges and then we would perform another iteration for smaller ranges.
################# Tune max_depth and min_child_weight ############## param_test1 = { 'max_depth':range(3,10,2), 'min_child_weight':range(1,6,2) } gsearch1 = GridSearchCV(estimator = XGBClassifier(learning_rate =0.1, n_estimators=100, max_depth=5, \ min_child_weight=3, gamma=0.3, subsample=0.8, colsample_bytree=0.8, \ objective= 'binary:logistic', nthread=8, scale_pos_weight=1, seed=27), \ param_grid = param_test1, scoring='log_loss',n_jobs=4,iid=False, cv=5)
Here, we see that the best combination is: max_depth = 5, min_child_weight=5, the best score is 0.4710678. We were going to go one step deeper and look for optimum values. We would search for values 1 above and below the optimum values.
param_test2 = { 'max_depth':[4,5,6], 'min_child_weight':[4,5,6] }
Here, we got best combination is: max_depth = 6, min_child_weight= 6, the best score is: 0.4706758. The score is slightly better. We can see that the score is better with higher values. We continue to increase the values of both parameters in the next search.
param_test3 = { 'max_depth':[6,7], 'min_child_weight':[6,7,8] }
Now, we see that the optimal values are: max_depth = 6, min_child_weight=7, the best score is 0.470626.
Then we performed similar approach to tune gamma, subsample and colsample_bytree, reg_alpha separately.
Grid search gamma:
| gamma | best value | score (log loss) |
|---|---|---|
| [0, 0.2, 0.4, 0.6, 0.8] | 0.4 | 0.4702224 |
| [0.3, 0.4, 0.5] | 0.4 | 0.4702224 |
Table 6: XGBoost Tune Parameter gamma
Grid search subsample and colsample_bytree:
| subsample | colsample_bytree | best combination | score |
|---|---|---|---|
| [0.6, 0.7, 0.8, 0.9] | [0.6, 0.7, 0.8, 0.9] |
|
0.4702224 |
| [0.75, 0.8, 0.85] | [0.75, 0.8, 0.85] |
|
0.4701294 |
Table 7: XGBoost Tune Parameters subsample & colsample_bytree
Grid search reg_alpha:
| reg_alpha | best value | score |
|---|---|---|
| [1e-5, 1e-2, 0.1, 1, 100] | 1e-5 | 0.4701294 |
| [1e-6, 1e-5, 1e-4] | 0.0001 | 0.47001335 |
Table 8: XGBoost Tune Parameter reg_alpha
Our last grid search ended with:
- colsample_bytree: 0.85
- gamma: 0.4
- max_depth: 6
- min_child_weight: 7
- reg_alpha: 0.0001
- subsample: 0.85
- Score: -0.47001335
We then fitted the model with the full training data and then got a much better training score -0.408851.
4.3.2. Logistic Regression
Parameter tuning in Logistic Regression is easier than the above tree-based models. We just did grid search on three parameters and performed a one-time search.
- penalty: Used to specify the norm used in the penalization (L1 or L2)
- fit_intercept: Specifies if a constant should be added to the decision function
- C: Inverse of regularization strength; smaller values specify stronger regularization.
logit = LogisticRegression(random_state=27, n_jobs = -1) para_grid = [{'penalty': ['l1', 'l2'], 'fit_intercept': [False, True], 'C':np.logspace(-5, 5, 10)}] para_search = GridSearchCV(logit, para_grid, scoring='log_loss', cv =5).fit(X_train, target_train)
We got the best combination:
- penalty = 'l1'
- C = 0.27825594022071259
- fit_intercept = True
This gave us the score -0.494125. We then fit the model with full training data and got a score 0.493337, which is not a big improvement.
4.4. Result
When we finally performed predictions with the test data using our three different models and submitted to Kaggle. Without any surprise based on the training score, XGB gave the best test score as well.
We also used model ensembling that combined our models to produce a hopefully improved results. Ensemble methods usually produces more accurate solutions than a single model would. We used a simple ensemble method: weighted voting. Using scikit-learn's VotingClassifier class, specific weights can be assigned to each classifier via the weights parameter. When weights are provided, the predicted class probabilities for each classifier are collected, multiplied by the classifier weight, and averaged.
Sample Code for voting ensembling with equal weight:
############ Try ensemble voting with average ########## ensemble_avg = VotingClassifier(estimators=[('lr', model_logit), ('xgb', model_xgb), ('extratree', model_extratree)], voting='soft', weights=[1,1,1]) ensemble_avg.fit(X_train, target_train)
We tried a few different combinations of selected models and weights, most of them didn't outscore the single XGBoost model. Only the combination of XGB + ExtraTrees with weight 3:2 slightly improved the score.
Table: Scores for different models
| Classifiers | Weight | Training Score | Test Score |
|---|---|---|---|
| Logistic Regression | N/A | 0.49334 | 0.49533 |
| ExtraTrees | N/A | 0.46891 | 0.47071 |
| XGB | N/A | 0.408851 | 0.46311 |
| Logistic Regression + ExtraTrees + XGB | 1:1:1 | 0.4372 | 0.46883 |
| Logistic Regression + ExtraTrees + XGB | 1:3:2 | 0.4249 | 0.46501 |
| XGB + ExtraTrees | 3:2 | 0.4134 | 0.46280 |
Table 9: XGBoost Tune Parameter reg_alpha
5. Conclusion
The BNP Paribas Cardif Claims Management competition is challenging due to its anonymous variables and high amount of missing data. During the competition, quite a few different methods were used for data cleaning, data imputation, and feature selections. Different machine learning classifiers such as Logistics Regression, ExtraTrees, XGBoost (Random Forest and GBM were being built but not yet completed) were used for training and predictions. Grid search technique was applied to fine tune our parameters in order to get the best model for each classifier. We also used model ensembling method "weighted voting", which slightly improve the score from one of combinations.
Many of the Kagglers in the competition have used the similar methods that we did. However some of them have higher scores than than us and the rest of the participant. We believe that what has made the difference is feature engineering. Though we applied some simple feature selection techniques such as tree-based feature importance and univariate feature selection, but obviously those were not enough to make a big improvement on the prediction accuracy. Therefore for our future work, we would like to look deeper on the data and do more feature engineering.
Reference
Jain, AArshay (2016, March 1). Complete Guide to Parameter Tuning in XGBoost (with codes in Python). Retrieved from http://www.analyticsvidhya.com/blog/2016/03/complete-guide-parameter-tuning-xgboost-with-codes-python/