Team DataUniversalis' Higgs Boson Kaggle Competition
Contributed by Shuo Zhang, Chuan Sun, Bin Fang, and Miaozhi Yu . They are currently in the NYC Data Science Academy 12 week full time Data Science Bootcamp program taking place between July 5th to September 23rd, 2016.
All R codes can be found here:
https://github.com/nycdatasci/bootcamp006_project/tree/master/Project4-MachineLearning/DataUniversalis/R_code
1. Introduction
Discovery of the long awaited Higgs boson was announced July 4, 2012 and confirmed six months later. But for physicists, the discovery of a new particle means the beginning of a long and difficult quest to measure its characteristics and determine if it fits the current model of nature. The ATLAS experiment has recently observed a signal of the Higgs boson decaying into two tau particles, but this decay is a small signal buried in background noise. The goal of the Higgs Boson Machine Learning Challenge is to explore the potential of advanced machine learning methods to improve the discovery significance of the experiment.
Using simulated data with features characterizing events detected by ATLAS, our task is to classify events into "tau tau decay of a Higgs boson" versus "background."
2. Work Flow
3. Pre-processing
3.1 Exploratory Data Analysis
3.1.1 Data Distribution Pattern
The previous research on the Higgs Boson revealed that the derived mass is an important indicator to determine its presence. By analyzing the distribution pattern of the variable DER_mass_MMC by label, we could see that the mass observations of “s” label has narrower distribution and higher peak value than “b” label. Mass observations of the two label groups have similar mean values.
3.1.2 Missingness
We noticed that only about a quarter of the dataset are complete cases. 11 variables out of 33 have missing records which are designated as -999. 10 out of the 11 variables are related to jets as well as the number of the jets.
By carefully examining the missingness through the jet numbers, it can be found that the variables aforementioned above corresponding to jet number 0 are all missing, while they are partially missing in jet number 1. There is no missingness detected in jet number 2.
PRI_jet_num = 0 PRI_jet_num = 1
PRI_jet_num = 2/3
3.1.3 Correlation Analysis
After imputing all -999 values for each jet number group, we can find high positive or negative correlations between a few variables, especially the variables with a prefix “DER”, and such correlations differ in different jet number groups. It can be inferred that these variables may derive from same origin.
PRI_jet_num = 0 PRI_jet_num = 1
PRI_jet_num = 2/3
3.1.4 Principal Component Analysis
With the help of principal component analysis (PCA), we can better understand importance and relationship among all variables. From the PCA scree plot it can be known that first component has contained most of information and the data dimension can be reduced down to 8 variables. The PCA correlation result shows that some variables with prefixes “PRI_jet” or “DER” have relatively higher loading values in the first two component PC1 and PC2 which may indicate the importance.
3.2 Imputation
After splitting data to 3 parts based on jet_number and dropping missing columns that have no physical meaning for each subset, there is still one important feature ("DER_mass_MMC") which has missing values for subsets with jet_number equal to 0 and 1. We applied KNN to impute missing values by choosing parameters k=sqrt(nrow(subset)) and distance=2.
4. Learning Algorithm Training
4.1 Overview of Machine Learning Methods
We have tried a number of different machine learning methods. Below is a quick review of the methods with a general description, their pros and cons.
4.2 Random Forests
Let us have a look at our first tree-based model, Random Forest(abbreviated as RF below). There are two parameters we need to tune for RF: ntree and mtry. Ntree controls how many trees to grow and mtry controls how many variables to draw each time. Due to the large size of the data, my first try was very ambitious: ntree = [2000,5000,8000] and mtry = [3,4,5,6]. However, it turned out that the computation was tremendously large and led to crush of R. So my second try was less ambitious with parameters ntree=[500,800,1000] and mtry = [3,4,5].
From the below graphs, we can see that mtry = 3 is the optimal parameter for each subset of data set.
However, the AUC curve looks abnormal because the graphs showed that the RF predicts every single observation correct. Why is that?
In our case, since weight is estimated from a probability space. In order to do prediction, RF model bisect on the probability space continuously until reach certain criterion (eg. no more than 5 observation in each sub-region) and use the mean of each sub-region to predict. However, our probability space is very sensitive and ntree = 1000 is too large thus making the RF too accurate.
What is more, by looking at only 3 variables (especially the traverse-mass variable), we can clearly tell whether the label is going to be a 'l' or a 'b'. So the first few split of the tree model is already sufficient to make the prediction. By making ntree equal 2 or 10, we can get a 'normal' graph. However, doing so is not very meaningful because it simply made a fine model less finer.
4.3 GBM
To improve RF, next step is to apply gradient boosting. There are 4 tuning parameters: interaction.depth, n.trees, shrinkage and n.minobsinnode. Interaction.depth controls maximum nodes per tree - number of splits it has to perform on a tree (starting from a single node). For example, interaction.depth = 1 : additive model and interaction.depth = 2 : two-way interactions. N.trees means the number of trees (the number of gradient boosting iteration), and increasing N reduces the error on training set, but setting it too high may lead to over-fitting.
Shrinkage is considered as a learning rate, which is used for reducing, or shrinking, the impact of each additional fitted base-learner (tree). It reduces the size of incremental steps and thus penalizes the importance of each consecutive iteration. The intuition behind this technique is that it is better to improve a model by taking many small steps than by taking fewer large steps. If one of the boosting iterations turns out to be erroneous, its negative impact can be easily corrected in subsequent steps.
N.minobsinnode controls the minimum number of observations in trees' terminal nodes, which is used in the tree building process by ignoring any splits that lead to nodes containing fewer than this number of training set instances. Imposing this limit helps to reduce variance in predictions at leaves. We can conclude the 4 tuning parameters are related with each other and change in one parameter has impact on the others.
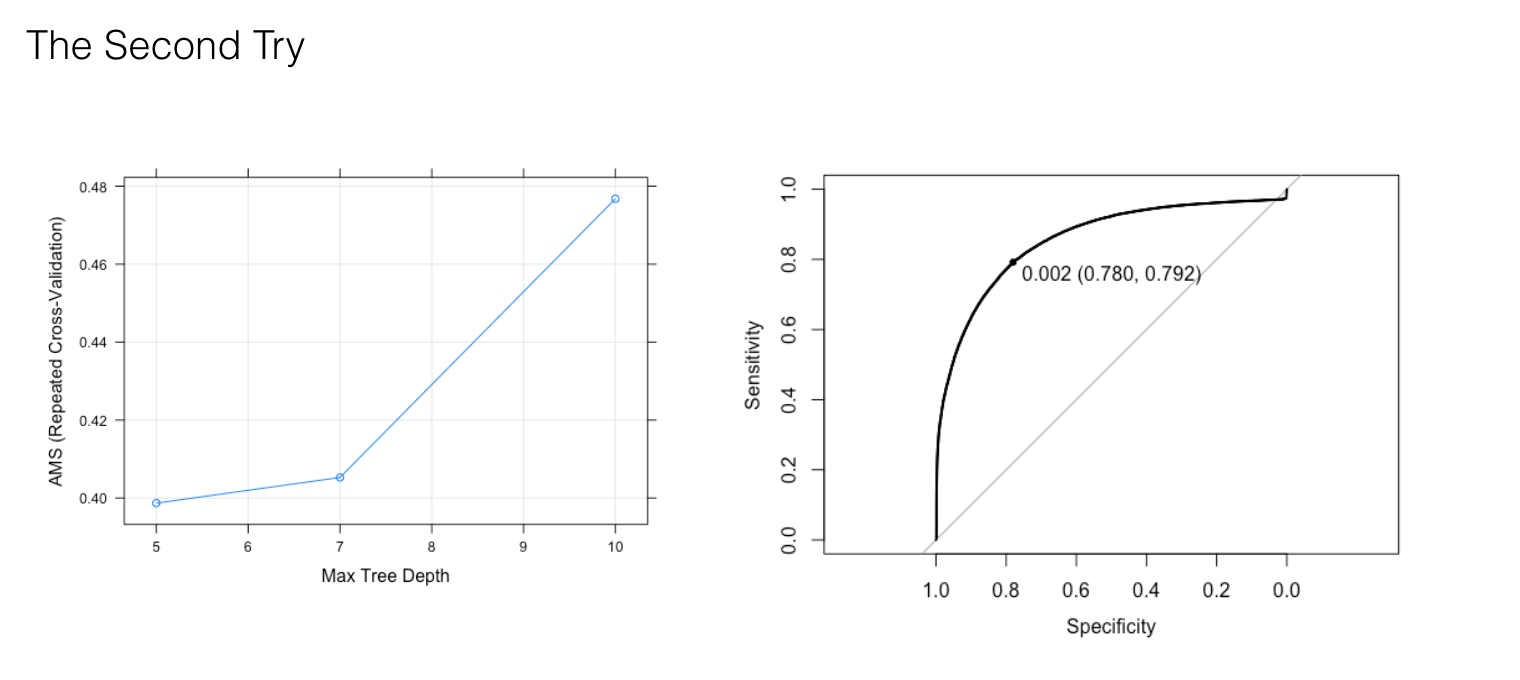
Let's take subset 1(jet_number equal to 1) for instance. Due to the large size of data and computational cost, our first try is to use small trees (200, 500, 800) with lower interaction depth (3, 4, 5), a wide range of shrinkage (0.1, 0.05, 0.01) and small values of n.minobsinnode (10, 50, 100). The best optimal parameters with the highest AMS value is 800 n.trees, 5 interaction.depth, 0.1 learning rate and 100 n.minobsinnode, which lead to AUC equal to 0.88(as shown in the graph). Therefore our second try is to fix n.trees and increase interaction.depth to (5,7,10).
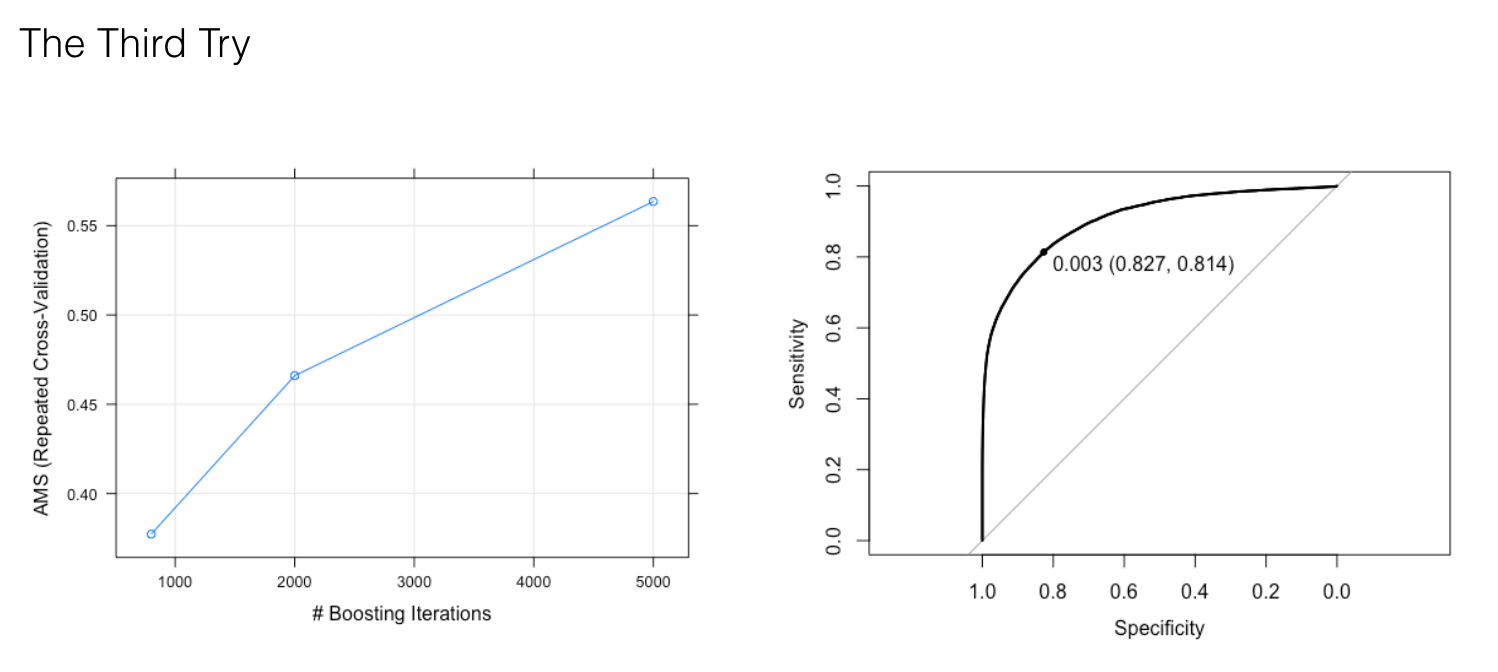
The best optimal parameters with the highest AMS is 10 interaction.depth, but it produces a lower AUC value equal to 0.86. The possible reason is overfitting by introducing too much interaction between the features. So our third try is to fix interaction.depth to 5, increase n.trees to (800, 2000, 5000) and lower shrinkage to 0.01 in order to reduce overfitting.
And 5000 trees results in a higher AUC value (0.91). So our last try is to increase n.trees to (5000, 7500, 10000) and fix interaction.depth and shrinkage to 5 and 0.01. Our best optical parameters (interaction.depth:5, shrinkage: 0.01, n.trees: 10000, n.minobsinnode: 100) give us the best result with AUC equal to 0.91.
The same process of tuning parameters is applied to the other 2 subsets.
For subset 0(jet_number equal to 0), the best tuning parameters are interaction.depth: 5, shrinkage: 0.01, n.trees: 10000, and n.minobsinnode: 100 and result in AUC equal to 0.91.
For subset 2(jet_number equal to 2 and 3), the best tuning parameters are interaction.depth: 5, shrinkage: 0.05, n.trees: 800, and n.minobsinnode: 100 and result in AUC equal to 0.91.
4.4 Neural Networks
The data set was split into three categories to do the neural network (NN) training. We tested several neural network tools under “Caret” library and chose “nnet”, as we thought one hidden layer would be appropriate for training. Two tuning parameters are available for controlling prediction accuracy: number of hidden units, which connects inputs and outputs and can feed into the output layer; weight decay, which is a parameter to control growing rate of the weights after each update.
We assumed that small weight decay and similar number of hidden units as number of variables would generate accurate predictions. We tried different numbers of hidden units ranging between 6-22, with an increment of 2, as well different number of decay weights between 0-0.1.
Below is the architecture of neural network when applies 1 hidden layer and 20 hidden nodes, where, blue lines indicate positive relationships while red lines indicate negative relationships.
Relative importance was acquired from neural network training using label as response variable. It can be noticed several variables with “DER” as prefix have higher importance, ranging 0.6-0.9, to label than the other variables. This shows consistency to the result of PCA analysis.
The result of NN parameter tuning on the data set when PRI_jet_num = 0 demonstrates that the NN training performance, which is reflected by AMS score is quite fluctuated as hidden node units increase. Great weight decay values (0.1, 0.01) have better AMS score than small values as well as clearer increasing trend of AMS can be observed. The AMS almost stays at 0 when weight decay is set as 0.0001. We found the best combination for training this dataset when hidden size of nodes = 10, weight decay = 0.01 and accuracy prediction = 82.54%.
After the first attempt of parameter tuning, it can be concluded that higher weight decay may positively contribute to AMS score and consequently a new range between 0.001-0.15 of weight decay was applied for NN parameter tuning on the data set when PRI_jet_num = 2/3. From the tuning result it can be found that most of AMS score line monotonically increase as more hidden units of nodes involved in NN training, only except when weight decay = 0.01.
Generally NN training performs better for weight decay = 0.1 or 0.15 than smaller weight decay values. The best set for training this data set is found when hidden size =22, weight decay = 0.1 and corresponding accuracy of prediction = 80.4%.
PRI_jet_num = 0
PRI_jet_num = 2/3
4.5 XGBoost
There are 6 parameters to tune for Xgboost.
| parameter | note | range | Our chosen value |
| nrounds | The number of rounds for boosting | [1,∞] | 20, 60, 100, 150, 200, 250 |
| eta [default=0.3] | step size shrinkage used in update to prevents overfitting. After each boosting step, we can directly get the weights of new features. and eta actually shrinks the feature weights to make the boosting process more conservative. | range: [0,1]. A value in between means we do not do full optimization in each step and reserve chance for future rounds, it helps prevent overfitting | 0.001, 0.003, 0.02, 0.05, 0.15 |
| gamma [default=0] | minimum loss reduction required to make a further partition on a leaf node of the tree. the larger, the more conservative the algorithm will be. | range: [0,∞] | 1, 5 |
| max_depth [default=6] | maximum depth of a tree, increase this value will make model more complex / likely to be overfitting. | range: [1,∞] | 7, 10, 15, 20 |
| min_child_weight [default=1] | minimum sum of instance weight(hessian) needed in a child. If the tree partition step results in a leaf node with the sum of instance weight less than min_child_weight, then the building process will give up further partitioning. In linear regression mode, this simply corresponds to minimum number of instances needed to be in each node. The larger, the more conservative the algorithm will be. | range: [0,∞] | 0.1 |
| colsample_bytree [default=1] | subsample ratio of columns when constructing each tree. | range: (0,1] | 0.7 |
Our tuning strategy for xgboost is as follows:
- Get an initial estimation of the range of each parameter, run scripts to get the best one, take a look at the “trend” from the saved graphs.
- Get rid of “bad” parameters that will not boost result, setup a new tuning grid, and run script again
- Repeat step 1 and step 2
5. Combining Predicted Probabilities
5.1 Analysis
We combined the training results from the three split datasets into one submission file, as illustrated in the hand-drawn picture below.
After the split, each subset corresponds to its own optimal tuned parameter. Then we were facing two strategies to combine the three vectors of probabilities:
- Strategy 1: Simply concatenate the three vectors (V0, V1, and V23) of predicted probabilities into one single vector V, then choose a uniform cutoff value C (i.e. C=15%) to map the V into a vector of labels containing 's' and 'b'. Finally submit this vector of labels to Kaggle to get a ranking.
- Strategy 2: Treat each subset differently. Each classifier of one subset has an optimal cutoff value based on the ROC curve. For example, the t0=0.35 on the ROC curve corresponds to the best threshold value of jet #0 . Using this threshold t0 as a cutoff value for all the predicted probabilities (eg., P01, P02, P03, etc), we can map the predicted probabilities to labels (signal 's' or background 'b') for all the testing observations that has jet number 0. As shown in graph below, P02 = 0.4 > t0 = 0.35, so P02 maps to 's'. Similarly, for the observations of jet #1, since P23 = 0.36 < t1 = 0.4, P12 maps to 'b'. We performed this procedure for jet #1 and jet #23 as well, concatenated the three vectors of mapped labels, and generated the submission file for Kaggle.
We tried both strategies multiple times and submitted our results to Kaggle. However, the rankings fluctuated around 1000 in the private leaderboard, which did not meet our expectations. Indeed, at the submission stage, it is possible for us to further tune a "magic" cutoff value, say, the value "C" in strategy 1 above, to obtain a great submission file that achieving relatively higher AMS score or ranking in the leaderboard, by submitting perhaps hundreds of times to Kaggle.
However, we didn't step into this direction, because we don't think repeatedly submitting results to Kaggle to achieve high ranking was an elegant solution. Trial and error may work for Kaggle, but remember in reality, we may only have one shot.
We sit back and started to figure out the root cause. Soon our team realized that, this combining process is actually mathematically flimsy.
5.2 Why splitting data by jet number is not a good idea?
Let's briefly revisit what made us want to split the dataset:
- This dataset happened to have 4 available jet numbers,
- Through EDA, we happened to find that the missingness of each jet number behaved differently
- We noticed someone in the Kaggle forum achieved high AMS score (~3.5) after splitting the data
- Splitting the data indeed led to customized imputation and faster parameter tuning
- We subjectively assumed that jet number was a very critical physics parameter
Let's again look at the examples in the hand-drawn graph. Although P03 = 0.36 in the subset jet #0 and P12 = 0.36 in the subset jet #1, the two probabilities 0.36 has totally different meaning:
- P03 = P(p03 | jet#=0) = 0.36
- P12 = P(p12 | jet#=1) = 0.36
In other words, the predicted probabilities in each subset were all conditional probabilities, a.k.a, conditioned on jet numbers. Thus, two identical probability values from two subsets mean different things, and they were not necessarily comparable with each other, because we had no prior knowledge about the internal distributions of each subset. Even if we do, do we know the signal distribution of the testing dataset? Not at all. The testing dataset could contain 15% observations with jet #0, or completely no jet #0.
To sum up, splitting the dataset based on jet number is indeed a possible way to achieve relative acceptable ranking or AMS score. However, based on our analysis, it also has drawbacks:
- Has the cost of finding another "magic" parameter at the final submitting stage
- Is mathematically shaky when combining predicted probabilities from each split
This means that splitting the dataset into 3 subsets was actually not a good strategy. Knowing this was very important since it made us think even deeper into our decision process as we navigated the project.
6. Takeaways
- Imputation turns out to be necessary after splitting the data (for EDA, feature importance, NN, RF)
- Try larger search grids for parameter tuning if possible (our grids were still too sparse)
- Use cloud computing for all team members not their own laptops (avoid crashing and uncertainties)
- Generally speaking, splitting data by jet number is not a good idea because it is mathematically shaky when combining predicted probabilities from each split, although relatively acceptable rankings in Kaggle can still be achieved