Hospital readmission prediction Data Analysis
The skills I demoed here can be learned through taking Data Science with Machine Learning bootcamp with NYC Data Science Academy.
Introduction
A goal of the Affordable Health Care Act is to increase the quality of hospital care in U.S hospitals. One specific area in which the Act tries to pressure hospitals to improve quality is by penalizing them for excessive readmissions for the same cause. Under the Affordable Care Act, the US Government's Centers for Medicare & Medicaid Services (CMS) created the Hospital Readmission Reduction Program to link Medicare and Medicaid payment data to the quality of hospital care to improve health quality for Americans. Fundamentally, payments to Inpatient Prospective Payment System (IPPS) hospitals depend on each hospital’s readmission rates, with, hospitals with poor readmission performance (excessive readmissions) are financially penalized through reduced payments.
A hospital in our scenario has a goal to reduce hospital readmission rates for diabetic patients. Both the hospital finance and clinical care teams are interested in how the data science team may help these departments reach this goal.
The measurements for hospital readmission are defined as:
- All-cause unplanned readmissions that happen within 30 days of discharge from the index (i.e., initial) admission.
- Patients who are readmitted to the same hospital, or another applicable acute care hospital for any reason.)
Our goal is to build a model which predicts whether a patient will be readmitted in under 30 days. A diabetic readmission reduction program intervention will use this model to target patients at high risk for readmission. The model will be evaluated by AUC and FPR values, lift table, sensitivity, and specificity, and finally will identify important features contributed to hospital readmission. The source code is available in our Github repository.
Data overview
Data were obtained from the UCI machine learning repository. This dataset contained clinical care at 130 US hospitals and integrated delivery networks from 1999 to 2008. There were 101,766 observations (71,518 unique patient Id). It included 50 features (13 numerical and 37 categorical features) such as race, gender, age, HbA1c test results, etc. This dataset satisfied the following criteria:
- It is a “diabetic” inpatient encounter (a hospital admission).
- Any kind of diabetes was entered into the system as a diagnosis.
- The length of stay was at least 1 day, and at most 14 days.
- Laboratory tests were performed during the encounter
- The medication was administered during the encounter

Figure 1 shows the distribution in the ratio of some features. In the age group figure, there are about 75% of patients in this dataset are older than 60 years old. In the gender plot, the female and male ratio is about 54% to 46%. As for the distribution of race, the Caucasian patient is the majority and follow by African-American patients in this data set. In the figure of the HbA1c test, more than 80% of patients didn’t conduct the HbA1c test during the encounter.

Figure 2 illustrates that about 90% of patients didn’t readmit within 30 days. Such imbalance dataset is challenging to make predictions. We will address this issue later.
Preprocessing
In this session, we will discuss how we conduct data processing and deal with imbalance data.
Missing data

There are eight features contained missing data, as shown in table 1. Since weight has nearly 97% of missingness, we decided to drop this feature. We also dropped the payer code feature because it was not considered irrelevant to the readmission rate. We retained the medical specialty and replaced those missing data with “Missing.” For the rest features with less than 5% of missingness, we added value “Missing” to account for those missing values.
Feature engineering

Three features were showing the diagnosis of each patient. The diagnosis features were represented by icd9 codes which contained more than 700 distinct values. We grouped those diagnosis codes into 9 different major groups to reduce the dimension of these features, shown in Fig. 3.
The raw dataset contained multiple visits for some patients, so those encounters cannot be considered as independent variables. Therefore, we only retained the first encounter (the smallest value of encounter Id) for each patient. The number of observations decreased from 101,766 to 71,518. Besides, we dropped the patients who were expired or discharge to hospice to avoid the bias of the model. The final number of observations was 69,973.
We dropped encounter and patient Id features because they are irrelevant to readmission rate. We transformed nominal features to string type to ensure they were treated as categorical features. We either dummified categorical features and applied label encoding before we fit the models.
Imbalance dataset

Figure 4 displays the distribution of readmission. The readmission rate is about 10% of the data. To deal with the issue of imbalance dataset, we applied the following three different techniques: oversample minority class, undersample majority class, synthetic minority oversampling technique (SMOTE).

Fig. 5 illustrates the procedure we applied to deal with imbalance dataset and the size of the training set after using each technique. We first applied the 80%/20% train-test split, and we then applied each sampling technique on the training data set. Therefore, we could train the models with balance dataset and evaluate our models with an unseen imbalanced dataset.
In the oversample minority class and SMOTE, we made the numbers of readmission as same as the numbers of non-readmission. The oversampling technique is just sampling with replacement of the minority class (readmission). SMOTE used the nearest neighbors algorithm to generate new and synthetic data we can use for training our model. The undersample majority techniques were to select the non-readmission observations without replacement randomly, and the sample size is the same as readmission observations.
Result and discussion
We trained five different models to make predictions as follows:
- Logistic Regression Model
- Logistic Model with Feature Transform From Gradient Boosting Model (GBM)
- Logistic Regression with Reduced Features
- Random Forest Model
- XGBoost Model (XGB)
We applied different data balancing techniques, as discussed in the previous session. We found that the logistic regression model predicted similar results for both over or under- sampling. Overall, logistic regression with SMOTE provided the best value on cross-validation.
The gradient boosting, random forest, and XGBoost models all show bias towards the no-readmission when we used models on test set when oversampling or SMOTE is used. However, these models performed well when we used under-sampling. Therefore, the results of GBM, random forest, and XGB we discuss below are using the under-sampling method. We will discuss the procedure in detail of the second and third models and compare the results with the other three models.
LR + GB Model
This Model first use half of the training set to fit a Gradient Boosting model, transform the features into a higher dimensional, sparse space, then the model was applied to the second half of the training set to extract features for the Logistic Regression model (Feature Transformation with Tress) as shown in Fig. 6. And Fig. 7 shows the top features. Table 2 & Table 3 are the results on the test set.




We extracted the top 5 features from the GB model (discharge_disposition_id, number_inpatient, num_dedications, diag_1_group, age )to the whole training data set to a Logistic Regression mode, achieved the same result as a full feature model. Table 4 & 5 are the results on the test set.

Model comparison

Figure 8 shows the Roc curves for the logistic regression, random forest, and XGBoost models. The performance of these models are similar to each other, but the XGBoost provide the highest AUC among these models.


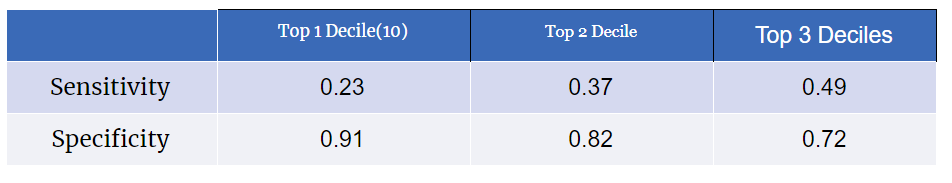
Table 6 shows the quantile groups predicted by the XGB model, and table 6 shows the value of sensitivity and specificity on different target groups. Compared to table 2 and 4, the XGB model provided the highest lift in the quantile 10. That’s being said, patients in the quantile 10 have the readmission rate 2.38 times higher than averaged patients. The XGB model also provided the highest value of sensitivity among different models. Therefore, we should use the XGB model to predict the readmission rate of future patients and filter out high-risk patients.

Figure 9
Fig. 9 demonstrates the feature importance of the XGB model. Discharge/disposition Id and the number of inpatient visits are the most important feature among different models. Age, diagnosis, and number of emergency also show up in Fig. 7. Those are the critical features to determine if the patient will readmit or not.

Even though we found that discharge/disposition Id is the most critical feature, we didn’t know which Id has the highest impact. Therefore, we used “get dummies” on logistic regression which provided important features in detail, as shown in table 8. The more information about the Id number can be found here. The top five highest coefficients are all from the discharge/disposition Id. The reference class for the discharge/disposition Id is “Discharged to home.” The top five Ids are all related to transferring to another medical facility. That means the patients who are discharged to another medical facility have much higher odd to readmit to hospital.
On the other hand, the features with the small odd ratio mean that those patients have less odd to readmit compared to patients in the reference class. These results can help us to find out how specific feature influence the readmission rate of each patient.
Summary
We used machine learning techniques to determine if diabetic patients readmitted within 30 days or not. In table 9, we list some key values to compare the performance of different models. Below are the summaries of our findings:
- Logistic regression performed well in both balanced (over or under-sampling) and unbalanced dataset. Also, the reduced feature model performs almost as good as the full model.
- XGBoost performed best among all the models we used.
- Tree-based models exhibit bias towards the majority when the dataset is unbalanced or over-sampling technique is used.
- Type of discharge and the number of inpatient visits are critical factors to determine readmission.

Future work
Since AUC is only 0.66, we can try different things to improve our model performance further. We could conduct more feature engineering building upon our current results. We could train some other models to see if any model provides higher AUC. Additionally, we could continue tuning hyperparameters on different models, such as XGB and gradient boosting, to obtain even better results. We would also like to investigate why Tree-based models bias towards the majority when the dataset is unbalanced or over-sampling method is used.
Applying model ensembling might also provide a more accurate prediction. Even though the discharge/disposition Id plays the most crucial role in the readmission rate, there are still many features not fully discussed. We will extract more information from our model results to provide better insights of which patients will have a high readmission rate. Our intention is that hospitals can reduce their readmission rates using the power of data science.

