Readmission Rates Increase: Developing a Reductions Program
Project GitHub | LinkedIn: Niki Moritz Hao-Wei Matthew Oren
The skills we demoed here can be learned through taking Data Science with Machine Learning bootcamp with NYC Data Science Academy.
Introduction
Readmission Reduction Program
The 3 main goals of the Affordable Health Care Act are to increase access to health insurance, protect patients from insurance companies and reduce healthcare costs. Through this, the act aims to improve the quality of the health care system in the US. Hospitals, as an integral part of this system, play a big role in achieving these goals. One specific way hospitals can improve quality and decrease costs is by reducing readmission rates.
Under the Affordable Care Act, Centers for Medicare and Medicaid Services has developed a hospital readmission reduction program that links payment to the quality of health care provided by hospitals and their readmission rates. The compensation to an Inpatient Expected Payment System (IPPS) hospital depends on the hospital's readmission rate. Hospitals with high readmission rates are subject to financial penalties in the form of reduced payments.
A condition patients tend to be readmitted for is diabetes, ranked seventh amongst the most deadly diseases in the US, afflicting nearly 9.4 percent of the population. In this project, we will develop a model to predict if a diabetes patient will be readmitted within 30 days. This will allow healthcare providers to take measures to prevent the patient from being readmitted.
To see our code, please click here.
Data
The data was provided by Virginia Commonwealth University, Center for Clinical and Translational Research. The dataset consists of 101,766 clinical care encounters of diabetic patients at 130 US hospitals and integrated delivery networks, collected between 1999 and 2008. It includes over 50 different information features about each encounter and patient.
Software/ Packages
For this project, we used Jupyter Notebook 4.4.0 version and python 3.7. The scikit-learn library was used for pre-processing as well as for all of machine learning models except for XGBoost.
Preprocessing
Exploratory Data Analysis
We started off by checking the type of data provided and discovered that there are 13 numerical features and 36 categorical features excluding the target variable (Readmission Status). 28 out of the 36 categorical features were ordinal while the remaining 8 were nominal.

The feature Readmission status has 3 categories but our goal is to predict the readmission within 30 days so this feature was converted to a binary variable with 1 representing readmittance within 30 days and 0 representing not readmitted or readmitted after 30 days. This left us with 89% (90,409) of the patients as 0 and around 11% (11,357) as 1.
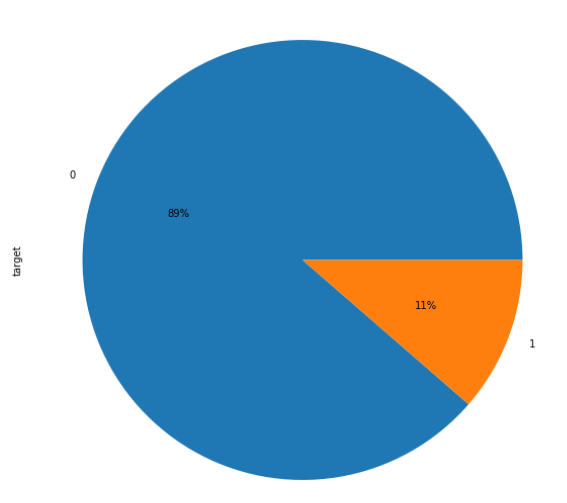
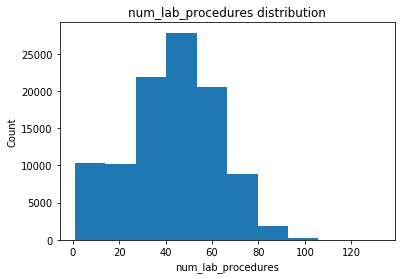
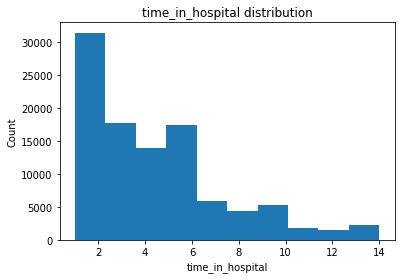
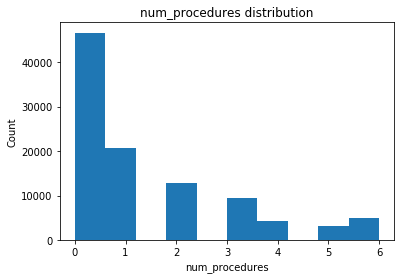
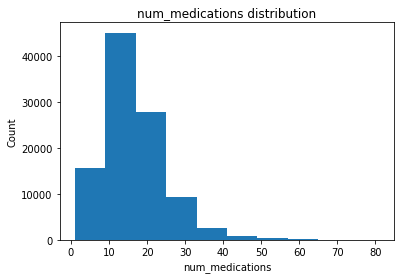
Distribution of some of the numerical features is shown below
Missingness & Imputation
Seven features had missing values represented by '?' which were converted to 'NaN'.
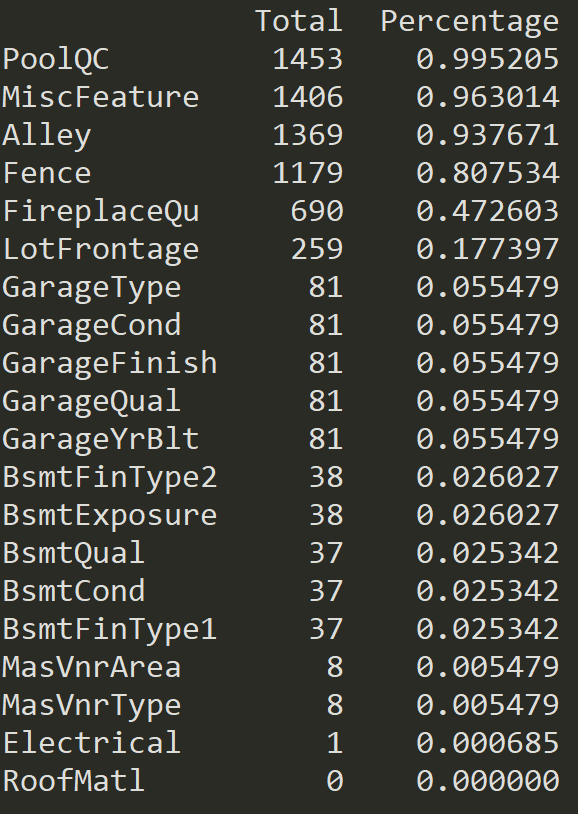
The weight variable was dropped as there were too many missing values. The missing values for race were filled with 'other' while missing values for diagnoses 1, 2, and 3 were converted to 0 corresponding to no diagnoses. For medical specialty, missing values were made into a new category 'No'. Missing values for payer code were converted to 'other'.
Feature Engineering
A number of discharge ids corresponded to a deceased patient. As these patients cannot be readmitted, these observations were dropped. There were also a number of diabetes medications that no patient was taking or had very little change. These medications were dropped from the data as well.
As there were a lot of id numbers for admission source id and discharge disposition id, many with only a few observations, we kept 9 ids with the most observations while all the other ones were combined to a category called 'other'. This reduced the number of categories to 10, making it easier to label encode and for the models to process.
Payer code was converted into 3 separate columns, self_pay, medicaid/Medicare, and coveredByIns all containing binary values. The observations with 'other' were represented by 0 in all three of these columns.
As we had 714 different type of diag ids, we matched the different ids to the corresponding ICD9 codes. We then grouped all the similar codes together to reduce the total number of codes to 18 different types. We also renamed diag_1, diag_2, ad diag_3 to f_diag, s_diag and t_diag. Once again this reduced the number of different categories for each feature, making it easier for the models to process.
After all this feature engineering was done, we used clustering to try to find similarities between observations. Plotting the number of clusters and the inertia showed the biggest improvement when using 2, 3 or 4 clusters. We made 3 new features with the values of the column corresponding to the cluster the observation belonged to. This grouped similar observations, potentially helping us to improve our predictions.


In trying to build a less complicated and more robust model, we also used feature importance from random forest to try to determine features that actually mattered. To do this, we added a feature with random binary values into the data and ran the feature importance.

The idea here is that we only choose the features that are more important than this random feature in the feature importance list. We do not want features added to our model that are less important than a feature with noise! In doing so we ended up with a data set with only 19 variables.
Data Prep
Once the data was ready, we kept 15% of the observations as a holdout set to test out our final models. The data was shuffled and split, keeping the ratio of the target variable the same. The remaining 85% of the data was split into a training set (70%) and a test set (30%), once again keeping the ratio of the target variable the same as the entire set.

As machine learning models require input features to be numerical values, we have to change the categorical variables to a format that the models can accept.
For logistic regression and SVM, the categorical variables were one hot encoded for the data. This dummyfication of a feature creates columns for each category of the feature containing binary values. This is essential because these 2 models can not process label encoded features as it treats them as numerical features instead of ordinal. Numerical features for these models were also standardized using StandardScaler from scikit-learn to address outliers and skewed distributions. StandardScaler was fit on the training data and then used to transform the training, test and holdout sets.

For the rest of the models, the categorical variables in the data were label encoded. This is a process where every category of a feature is mapped to a numerical value, resulting in a feature that we can process through our models.
Modeling
Once the data was prepped, we were ready to run different classification models to see which one was best suited to our particular problem. We chose to use logistic regression, random forest, XGBoost, gradient boosting, and support vector machines. Our goal was to find the model with the highest AUC.
As our data set was imbalanced, with the negative class (0) dominating with 89% of the observations, it was important that we addressed this issue before running the models. With such an imbalanced data set, the models tend to predict every observation as the dominant class. The models can predict with 89% accuracy but without a single true positive, making the models useless.
This problem can be addressed by a few methods. The minority class can be 'oversampled' by duplicating the positive observations until the number of observations for both classes is equal. We can also 'undersample' by only taking a random sample of observations of the majority class equal to that of the minority class so that we have a balanced data set. Scikit-learn models also have an in-built mechanism to balance a data set and this is the option we chose for our models. One such example is shown below.

To tune the hyperparameters for our models, we ran RandomizedSearchCV, a cross-validation method in scikit-learn. As RandomizedSearchCV is considerably faster than GridSearchCV and gives similar results we used it as the primary cross-validation method for some models. However, for some models, we used it as a first step to get an estimate of the hyperparameters before running GridSearchCV with a narrower range. Sample code for one of the models is shown below.
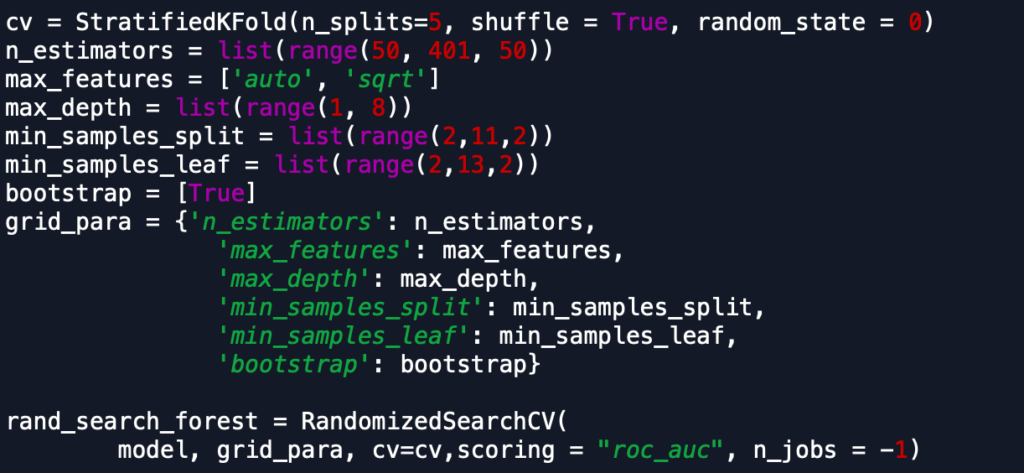
We used StratifiedKFold to ensure that our class ratio was the same for the folds used in cross-validation. We also used AUC as our scoring method as using accuracy for classification in an unbalanced data set is not ideal. As mentioned before, this returned hyperparameters that predicted all the observations as the majority class. During tuning the parameters, our goal was to try to get our models to have a high AUC and recall but also have high precision.
We also opted to run the random forest model with the reduced data set consisting of only the important features. However, this model performed worse than the full data set so we did not use this method with any of the other models.
Results
After the cross-validation and hyperparameter tuning, all the trained models were used to predict readmission for the test set. As expected, the AUC and recall score for the models were slightly lower than the scores for the training set. While the AUC for all the models were similar, there was more variation in the recall scores between models. Logistic regression had the lowest recall score while XGBoost performed the best out of the models.

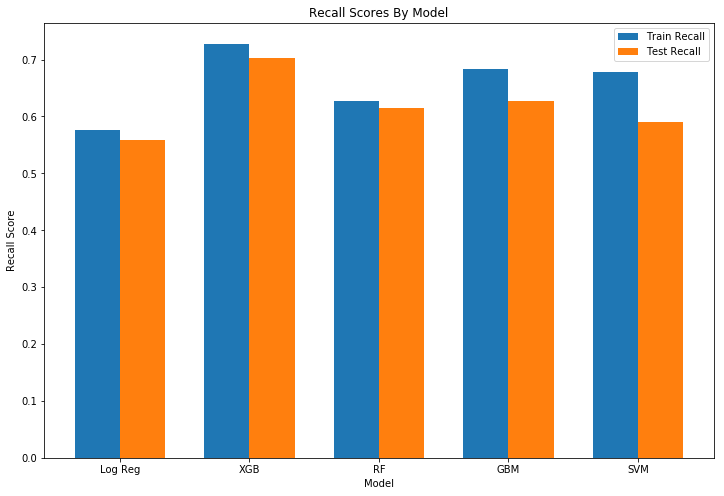

Finally, we tested the models on the holdout set to see how well they performed.

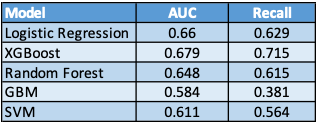
However, the precision of all the models was very low, something which is common in the medical field. In trying to improve recall, there were a lot more false positives being predicted by our models. To improve precision, we decided to focus on patients with a higher risk of readmission within 30 days predicted by our models.
To do this we used our models to predict the probability of readmission within 30 days for the train/test data set. Based on these probabilities, the data was split into percentiles with 844 observations in each. For each percentile, we got the range of the probability of being readmitted, as well as the precision. All the models had a better precision in the higher percentiles, but this score dropped as the percentile decreased.

To make a decision on the best model, we chose to look at a combination of the AUC/ recall score as well as how well each model did at predicting readmissions for the high-risk percentiles. Based on these criteria, we chose XGBoost as our final model as it had the highest AUC/ recall and also had the highest precision for high-risk patients.
Feature Importance - XGBoost

The feature importance of the model gave us an idea of the factors that were potentially driving patients to be readmitted within 30 days. The most important feature for our selected model, as well as all other models, was the number of inpatient visits. The higher the number of inpatient visits, the higher the probability of the patient being readmitted. This same relationship could also be seen with number of emergency visits, number of diagnoses and time in hospital which were the 3 other numerical features in the top 10.
Discharge ID was also an important feature for all the models. After taking a detailed look at the different discharge IDs, one, in particular, stood out. For ID # 22, which referred to a patient being transferred to a rehab facility, 27% of the patients were readmitted, compared to the general 10% of the sample. This ID showed up only 1% of the time for patients who weren’t readmitted but jumped to 5% for the ones who were readmitted.
As expected, age was an important factor for this model. The patient being on diabetes medication also affected the readmission rates as did metformin, a common diabetes medication.
Model application for hospitals
This project was designed to help hospitals decrease readmission rates for diabetic patients. With the proposed model, hospitals can target patients in high-risk percentiles. Not only are these patients at higher risk of being readmitted, the model precision is considerably better for the higher percentiles which means hospitals can efficiently use their resources to reduce readmission rates. Medical facilities can take precautionary measures with these patients during their initial admission or at the time of discharge or schedule a follow-up visit to check their progress.
This allows hospitals to provide a better quality of healthcare to their patients and also reduce the readmission rates. This reduction can help hospitals avoid penalties that are incurred for high readmission rates, leading to an improved bottom line.
Future Work
Given more time, we could run stacked models to improve the AUC and recall. The current models can be improved further through finer tuning of the hyperparameters. We can also dive deeper into the most important features of the models to see which particular categories of the features are affecting the classification. We can also try to change the classification threshold for some models to see if they improve the performance, especially reducing the false positives to improve the precision score.